Mapping in R: Street Trees in Cambridge-Somerville
Eric Robsky Huntley, PhD (They/them)
- Lecturer in Urban Science and Planning
- ehuntley@mit.edu
Tutorial Information
- Module 5 in Spatial Data Science.
Methods
Tools
Read Data
We begin by reading the tree census data I’ve provided. These are two
separate shapefiles, so we will read them independently using the
st_read function. Both contain quite a bit of potentially
useful information about each surveyed tree—especially Cambridge, whch
provides a full 60 attributes! However, for our simple analysis today,
we’re going to satisfy ourselves with each tree’s point location, stored
in the ‘geometry’ column of each layer.
require('sf')
require('dplyr')
require('tibble')
cambridge_trees <- st_read('data/cambridge_trees.shp') %>%
select(c('geometry'))somerville_trees <- st_read('data/somerville_trees.shp') %>%
select(c('geometry'))With this single column selected, we can elect to combine the two
using the rbind function to bind the sf
objects by row (hence,r). Plotting the trees, we see a
street canopy that hews quite closely to what we know to be
Cambridge-Somerville’s built form.
trees <- rbind(cambridge_trees, somerville_trees)
plot(trees, pch='.', col='green')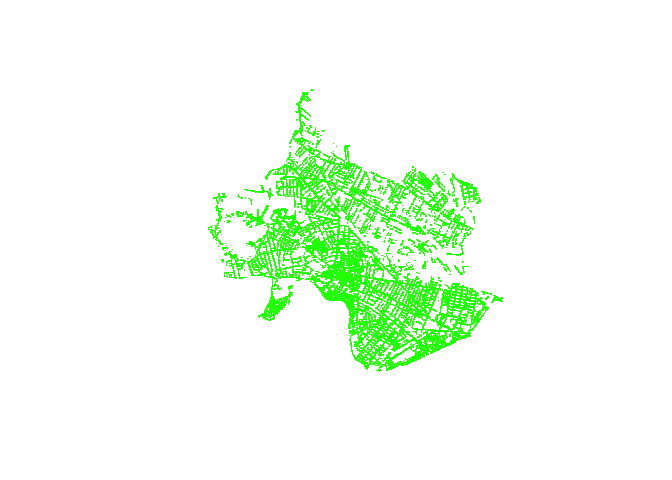
Generate Hex Grid
We now want to generate a grid of consistently-sized cells over which to evaluate the density of street trees in Cambridge-Somerville. Specifically, we want to generate a hexagonal grid.
Geographers love hexagons! This is because the distance from the center of a hexagon to its edge is approximately constant. This means that points included in a hexagon are going to be much closer to those that are within a threshold distance. This is not true for a square grid! For a square, $ d_{orthoganal} = d_{diagonal} $, where $ d_{orthogonal} $ is the distance from the center of the square and $ d_{diagonal} $ is the distance from the center of the square to any given corner.
We can use the st_make_grid function to generate a
regular hexagonal grid of half-mile cells. square=FALSE
instructs the function to generate hexagons. Finally, we convert the
simple feature collection (sfc) geometries that are created
to an sf object using st_sf and create an id
column called hex_id using
rowid_to_column().
hex <- st_make_grid(trees, cellsize=2640, square=FALSE) %>%
st_sf() %>%
rowid_to_column('hex_id')
plot(hex)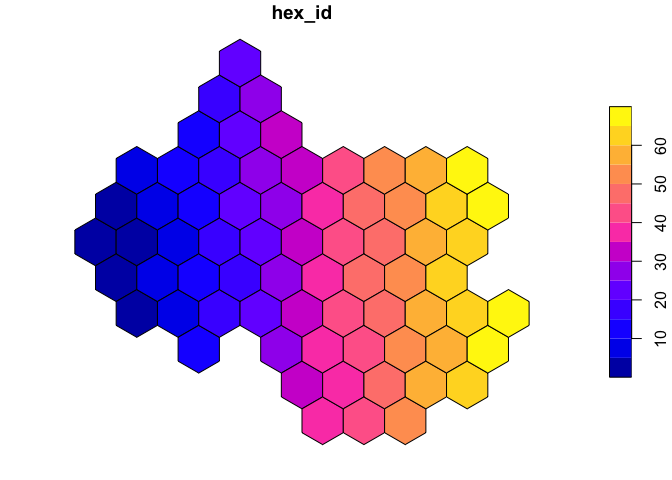
You can, of course, use different cell sizes; in general, you’ll find that larger cell sizes lead to more preditable behavior (tighter clustering around the sample mean, etc.), but that you lose spatial granularity as the cell size increases.
Count Points in Polygons
Now that we’ve created a grid, we can count the number of trees in
each polygon quite simply. We perform a spatial join between the trees
and the hexbins, using a st_within criterion. This means
that trees will be joined to the hexagons within which they fall. We can
plot the results and see that each tree has taken on the
hex_id of the hexagon within which it falls.
trees_in_hex <- st_join(trees, hex, join=st_within)
plot(trees_in_hex, pch='.')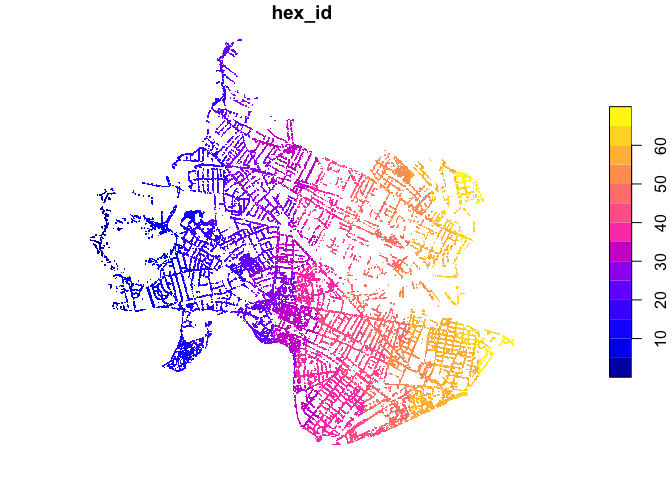
Ultimately, though, we’re not interested in the tree geometries, but
in the count per hexagon cell. As such, let’s remove the geometry from
the joined table and count the number of trees falling within each cell
by grouping on the hex_id. We can do all of this in a
single step using dplyr syntax like so…
trees_in_hex <- st_join(trees, hex, join=st_within) %>%
# Remove geometry.
st_set_geometry(NULL) %>%
# County the number of rows per hex_id and
# Store this number in a field named 'trees'.
count(name='trees', hex_id)
head(trees_in_hex)## # A tibble: 6 x 2
## hex_id trees
## <int> <int>
## 1 2 11
## 2 3 131
## 3 4 49
## 4 5 253
## 5 6 14
## 6 7 607Finally, we can use left_join to join the resulting
trees_in_hex table to the hex table, retaining
even those cells with no trees. Hexagon cells without any street trees
will then have NA values. Let’s replace these with
0s, because we know that these have no recorded street
trees—it is not only that values are unknown or missing.
hex <- hex %>%
left_join(trees_in_hex, by = 'hex_id') %>%
replace(is.na(.), 0)
plot(hex['trees'])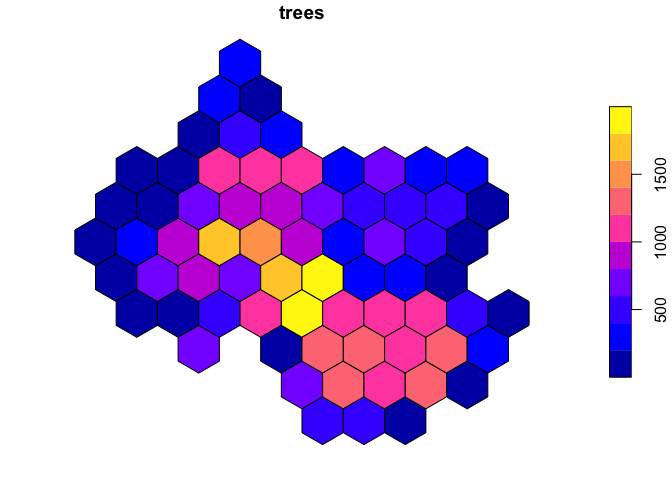
Create a Simple Map
We now have an analytically useful, if simple, dataset! However,
those plot functions are quickly reaching the outer edges
of their usefulness. Geographic analysis is often an exercise in
discerning the relationship between phenomenon and context—said simply,
a basemap would be a useful thing to have! Interacting with our maps
also becomes quite useful as we begin to explore Cambridge-Somerville’s
urban canopy.
For these reasons, the R wrapper for Leaflet is quite a useful thing. Leaflet is one of the most common and usable interative mapping libraries around; the fact that there is now a package that allows us to interface with its functionality without ever leaving R is marvelous. And it’s quite simple to use! Let’s make a first map.
We’ll create a new leaflet object, and—again, using the
piping syntax—set its view such that it is centered on
Cambridge-Somerville, and such that its view is set to a moderate
13. Map tiles (and therefore web maps) use standard zoom
levels from 0-20, where 0 is the least-zoomed-in and 20 is the most.
If we assign our map to a variable name, we can print it by simply invoking the variable name.
require('leaflet')
map <- leaflet() %>%
setView(lng = -71.111408, lat = 42.38172, zoom = 13) %>%
addTiles()
map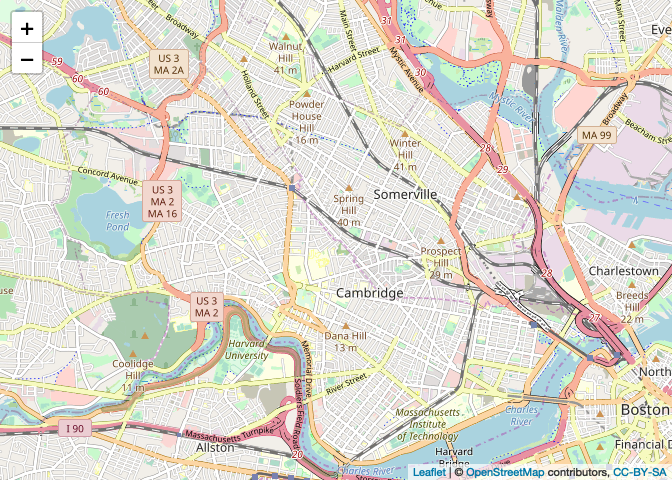
By default, leaflet will use an OpenStreetMap-styled
basemap. Nothing wrong with that! After all, most (non-Google, Apple, or
Bing) web map tilesets are just styled OpenStreetMap data. But if you
want a slightly more stylized basemap, R’s leaflet provides
a providers object that lists many available basemap
providers. These line up with those made available through the Leaflet
providers interface, if you want a slightly more visible way to
browse.
providers[1:5]## $OpenStreetMap
## [1] "OpenStreetMap"
##
## $OpenStreetMap.Mapnik
## [1] "OpenStreetMap.Mapnik"
##
## $OpenStreetMap.DE
## [1] "OpenStreetMap.DE"
##
## $OpenStreetMap.CH
## [1] "OpenStreetMap.CH"
##
## $OpenStreetMap.France
## [1] "OpenStreetMap.France"Once you’ve landed on one you like, you can load it using the
addProviderTiles function, with the associated tileset
name.
map <- leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite")
map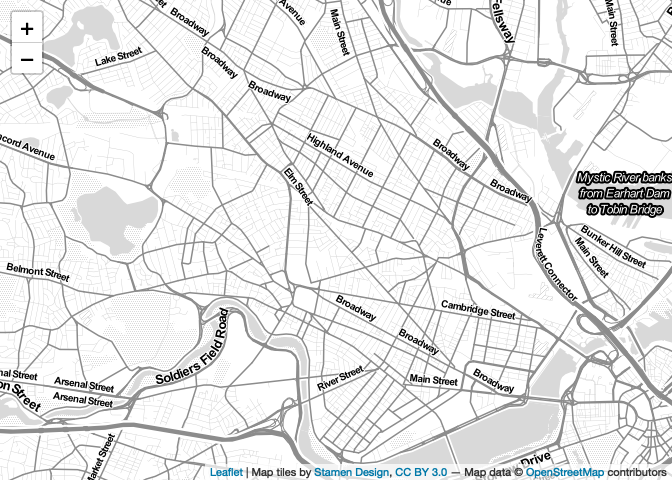
At this point, we’ve included a basemap, but we have yet to include
our data. Let’s change that! We begin by passing our hexbins through the
leaflet object, and add the geometries by using the
addPolygons() function. Notice that the
leaflet object knows to create polygons based on the
dataset piped to the constructor.
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerBackground") %>%
addPolygons()## Warning: sf layer is not long-lat data
## Warning: sf layer has inconsistent datum (+proj=lcc +lat_1=42.68333333333333 +lat_2=41.71666666666667 +lat_0=41 +lon_0=-71.5 +x_0=200000.0001016002 +y_0=750000 +ellps=GRS80 +towgs84=0,0,0,0,0,0,0 +units=us-ft +no_defs ).
## Need '+proj=longlat +datum=WGS84'map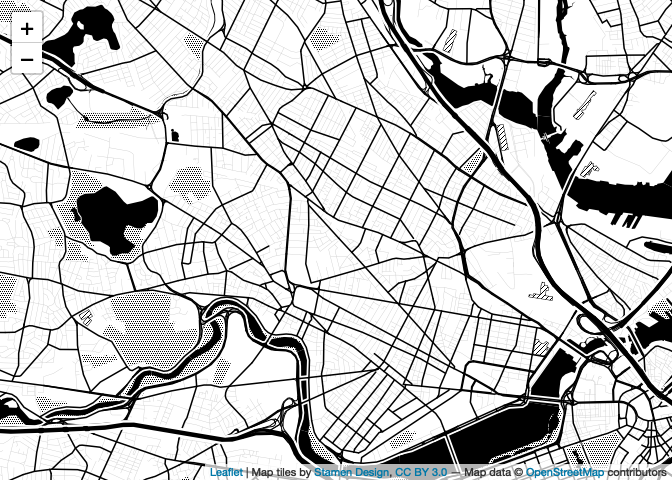
Oh no! Why are we seeing this
error?sf layer is not long-lat This is because leaflet
expects data that is stored in the WGS 84 coordinate reference system
(projection). As demonstrated last week, we can reproject data quite
simply using st_transform, and an EPSG code (in this case EPSG:4326).
hex <- st_transform(hex, 4326)
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerBackground") %>%
addPolygons()
map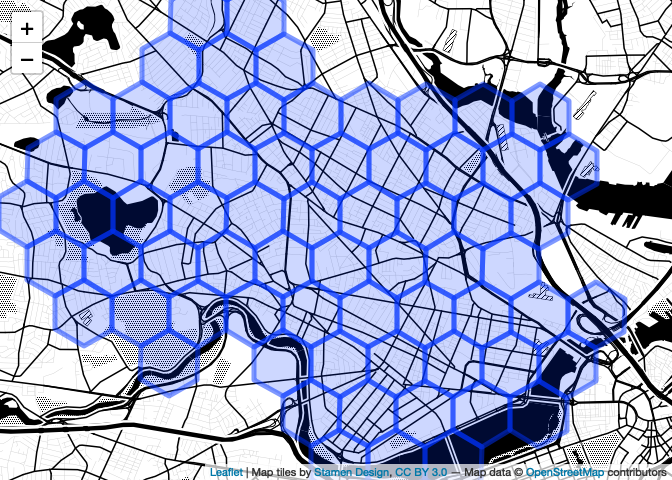
Better! Note that if you don’t like the ‘pipe the data to the leaflet constructor’ syntax I’m using, this is a perfectly fine alternative, which does the same thing and perhaps the reduces the ambiguity.
map <- leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerBackground") %>%
addPolygons(data = hex)
map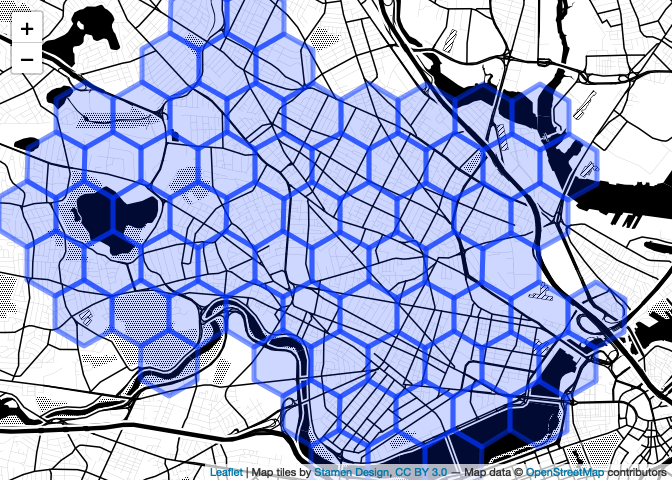
Choropleth!
We can extend this code ever-so-slightly to produce a choropleth map
that visualizes our street tree counts. We do this using the
ColorBin function to create a palette function that will
assign a color to each hexagonal cell based on its data value. We pass a
color ramp, a domain (i.e., a range of all possible values), and a
number of bins. By default, colorBin uses an equal-interval
classification scheme.
palette <- colorBin('Greens', domain = hex$trees, bins = 5)
palette(c(400,800, 1250))## [1] "#EDF8E9" "#BAE4B3" "#74C476"Greens here refers to the Greens
Colorbrewer color ramp… to see all available RColorBrewer color ramps,
simply invoke RColorBrewer’s
display.brewer.all().
require('RColorBrewer')
display.brewer.all()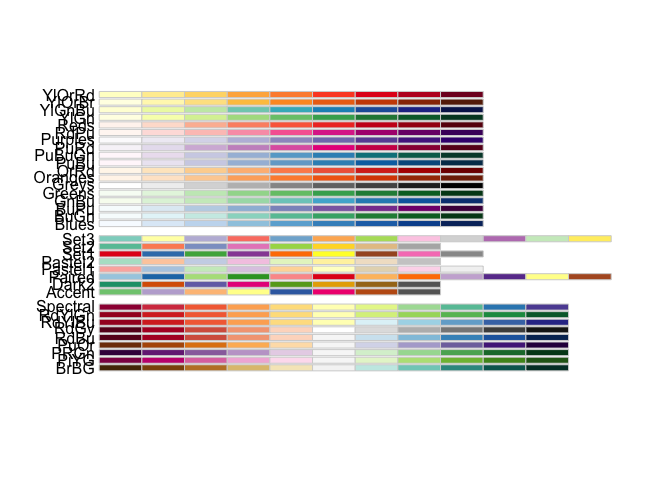
With this new palette in hand, we can modify our
addPolygons function call as below. Note that I also begin
passing in other style parameters—fillOpacity,
opacity (stroke opacity), weight (stroke
weight), and color (stroke color).
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
# for each row, assign a fill color based on the
# value returned by the palette() function
addPolygons(
fillColor = ~palette(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white'
)
map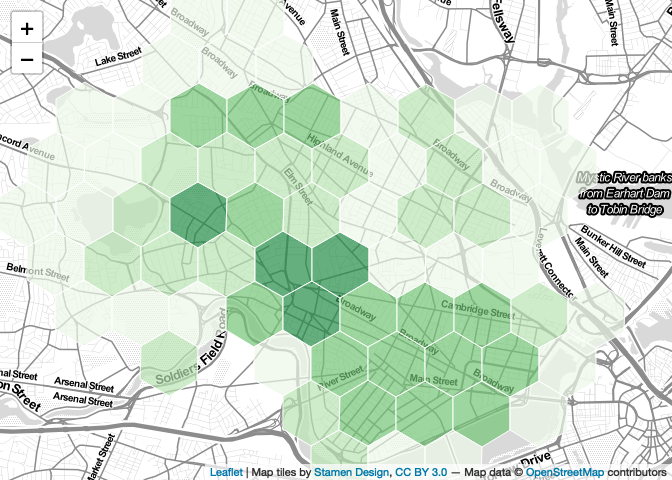
Control Your Class Breaks…
We know, though, that class breaks can have quite a dramatic effect
on the interpreting of visualized data. We don’t necessarily want to
simply accept an equal intervals scheme and call it a day! It’s good,
then, that Jenks natural breaks, quantile breaks, and many others are
readily available using the classInt package, which
interfaces quite neatly with the ColorBin function.
For example, if we wanted quantiles, broken across seven classes, we
would first create the quantile breaks using the
classIntervals function with a value of
quantile passed to thestyle parameter…
require('classInt')
quantiles <- classIntervals(hex$trees, n=7, style='quantile')
print(quantiles)## style: quantile
## one of 74,974,368 possible partitions of this variable into 7 classes
## [0,129.2857) [129.2857,223.5714) [223.5714,481.2857) [481.2857,660.7143) [660.7143,923.4286) [923.4286,1186.286) [1186.286,1974]
## 10 10 10 9 10 10 10print(quantiles$brks)## [1] 0.0000 129.2857 223.5714 481.2857 660.7143 923.4286 1186.2857 1974.0000Note that this function returns a few different variables—we’re
interested in its brks, which is a list of values that form
break points for our classification scheme. We can then pass this to the
bins parameter of colorBin.
palette_quantiles <- colorBin('Greens', domain = hex$trees, bins=quantiles$brks, pretty=FALSE)We can then modify our map to use this palette instead of the default palette.
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
# for each row, assign a fill color based on the
# value returned by the palette() function
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white'
)
map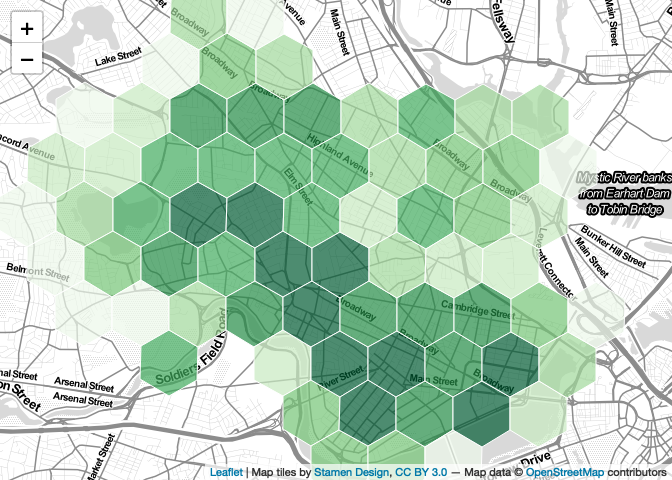
You could do similar exercises for Jenks natural breaks, equal
interval breaks, or any of the other classification schemes available in
classIntervals—the list is quite long! See
?classIntervals.
# Jenks
jenks <- classIntervals(hex$trees, n=7, style='jenks')
palette_jenks <- colorBin('Greens', domain = hex$trees, bins=jenks$brks, pretty=FALSE)
# Equal Interval
equal <- classIntervals(hex$trees, n=7, style='equal')
palette_equal <- colorBin('Greens', domain = hex$trees, bins=equal$brks, pretty=FALSE)Add a Legend and Scale Bar
Leaflet makes it quite simple to automatically generate a legend for
your map. It interprets the same palette functions that
we’ve been using to determine where class breaks are, and to which
colors they correspond. Furthermore, it includes a
labFormat parameter that takes a labelFormat
function as input, which gives you access to a number of simple ways to
control the legend display. Here, I round my legend breaks to 0 digits
after the decimal.
Finally, we can add a dynamic scale bar using the
addScaleBar function, providing a only position.
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
# for each row, assign a fill color based on the
# value returned by the palette() function
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white'
) %>%
addLegend(
title = "Number of Street Trees",
pal = palette_quantiles,
opacity = 0.7,
values = trees,
labFormat = labelFormat(
digits = 0
)
) %>%
addScaleBar(position='bottomleft')
map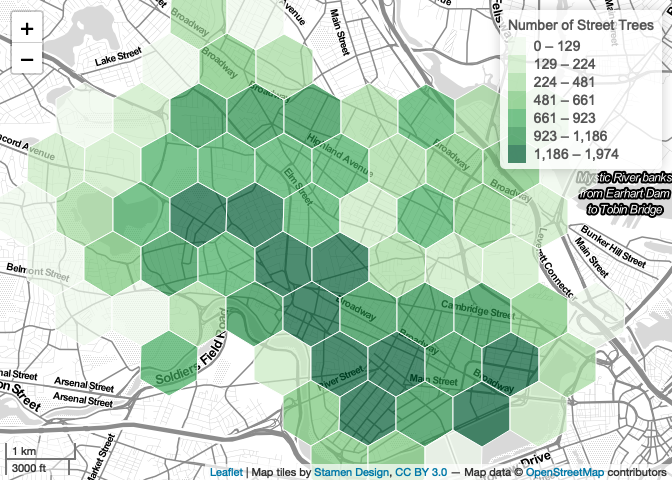
Make it a Bit More Interactive…
Leaflet also gives you access to a number of convenient functions that allow us to introduce simple behaviors that respond to user interactions. For example, we can introduce a highlight interaction that changes the style of a polygon as the user mouses over. We increase the line weight, and bring the feature to the front.
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white',
# Highlight behavior
highlight = highlightOptions(
weight = 3,
bringToFront = TRUE),
) %>%
addLegend(
title = "Number of Street Trees",
pal = palette_quantiles,
opacity = 0.7,
values = trees,
labFormat = labelFormat(
digits = 0
)
) %>%
addScaleBar(position='bottomleft')
map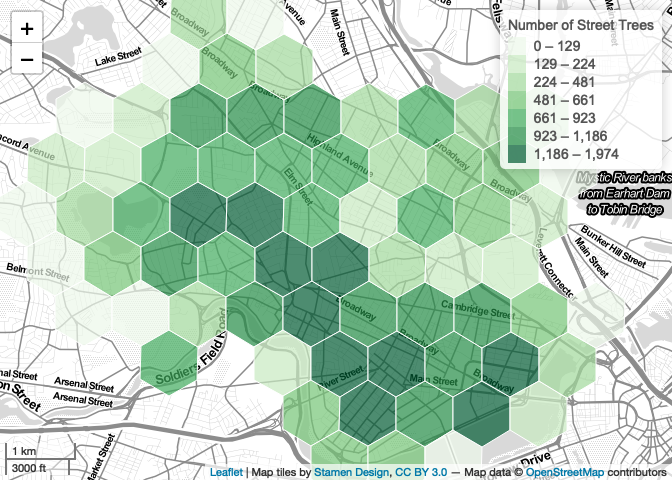
Simple interactivity tweaks like this encourage the map viewer to play, interact, and explore! But if we’re going to entice their further attention like this, we should at least give them a bit more information for their troubles… in addition to highlighing we can also add pop-up labels that print the tree count and hexagon id.
First, we create a list of labels—one for each hexagon! This code
looks complicated, but let’s break it down… for each row, we use the
stringr package’s str_glue function to create
a string that includes both the value of trees and
hex_id. In this string, we use HTML markup to indicate that
we want XXX trees to be bold
(<strong></strong>). Finally, we use
lapply to mark each item in the list as HTML. This means
that the HTML markup will not be displayed as text, but interpreted as
HTML.
require('stringr')
labels <- str_glue("<strong>{hex$trees} trees</strong> in hexagon {hex$hex_id}") %>%
lapply(htmltools::HTML)We can then add this to our map like so:
map <- hex %>%
leaflet() %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
addScaleBar(position='bottomleft') %>%
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white',
highlight = highlightOptions(
weight = 3,
bringToFront = TRUE),
# Add labels here.
label = labels
) %>%
addLegend(
title = "Number of Street Trees",
pal = palette_quantiles,
opacity = 0.7,
values = trees,
labFormat = labelFormat(
digits = 0
)
)
map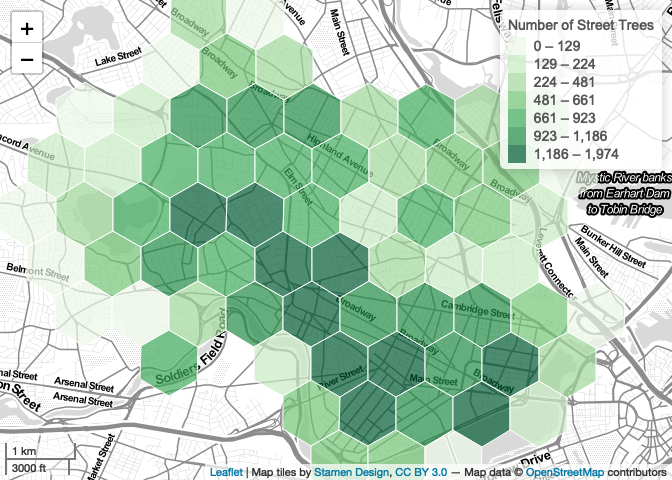
# mapshot(m, url="output.html")That’s it! We now have a map that displays additional information with user interaction… not too bad for a few lines of code, without ever leaving R!
Tidying Up
However, as we’ve made our map more interactive, you may have noticed
some annoying features; it’s possible for the user to zoom out far
further than would be useful in our application. The same can be said
for panning: the user can pan around the world, potentially becoming
quite lost. Finally, if you’re a person who’s done quite a bit of
programming, it might be making you itch that we’ve hard-coded the
setView latitude and longitude. Let’s deal with each one of
these things in turn. First: setting a minimum zoom.
Set a Minimum Zoom
We can set a minimum zoom using the minZoom parameter of
the leafletOptions function. When a minimum zoom is
specified, the user will not be able to zoom out further than the
minimum. Here, we set it to 12—one ‘step’ out from the
default of 13. Simple enough!
map <- hex %>%
leaflet(
options = leafletOptions(
minZoom = 12
)
) %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
addScaleBar(position='bottomleft') %>%
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white',
highlight = highlightOptions(
weight = 3,
bringToFront = TRUE),
# Add labels here.
label = labels
) %>%
addLegend(
title = "Number of Street Trees",
pal = palette_quantiles,
opacity = 0.7,
values = trees,
labFormat = labelFormat(
digits = 0
)
)
map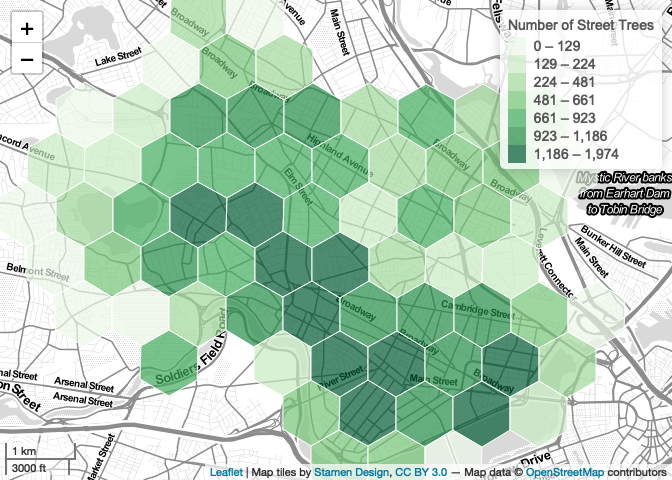
Bound the Map
In addition to ensuring that the user doesn’t have the ability to accidentally zoom out much farther than is useful, we may also want to put a fence around their horizontal panning. This is very useful—one of the more frustrating things when interacting with a map is dragging around, trying to find your starting point after an vigorous click-and-drag sends you sliding off into the world.
We set this using the setMaxBounds function, which takes
four parameters: the latitude and longitude of the top-left corner of a
bounding box, and the same for the bottom-right. We can programmatically
generate these by simply using the bounding box of the hexagons, which
are conveniently accessible to use using the st_bbox
function.
bbox <- st_bbox(hex)
print(bbox)## xmin ymin xmax ymax
## -71.16930 42.34927 -71.06187 42.42051We want to pass only the values, excluding their names, to the
parameters of the setMaxBounds function—to the R user, this
means using the double-bracket! Double-brackets allow you to access only
values, stripping away names.
map <- hex %>%
leaflet(
options = leafletOptions(
minZoom = 12
)
) %>%
setMaxBounds(
lng1 = bbox[['xmin']],
lat1 = bbox[['ymin']],
lng2 = bbox[['xmax']],
lat2 = bbox[['ymax']]
) %>%
setView(-71.111408, 42.38172, 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white',
highlight = highlightOptions(
weight = 3,
bringToFront = TRUE),
label = labels
) %>%
addLegend(
title = "Number of Street Trees",
pal = palette_quantiles,
opacity = 0.7,
values = trees,
labFormat = labelFormat(
digits = 0
)
)
map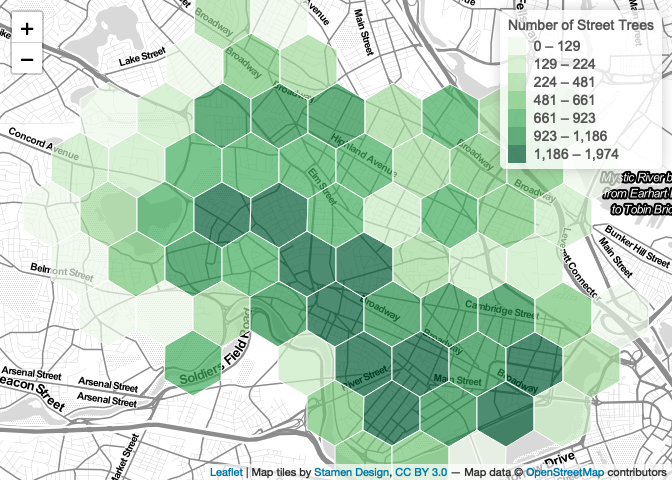
Setting MaxBounds makes interacting with and exploring your map much less frustrating for users.
Programmatically Center the Map
If you’re a person who does much programming, you may be bristling at
the fact that we’re manually passing in latitudes and longitudes to the
setView function. Instead, you might ask, isn’t there a way
to set this programmatically based on the center of our dataset? Yes,
dear reader, there is.
Let’s calculate the center of our dataset. First, we perform a union, which dissolves the edges between adjacent polygons, producing a single large polygon…
plot(st_union(hex))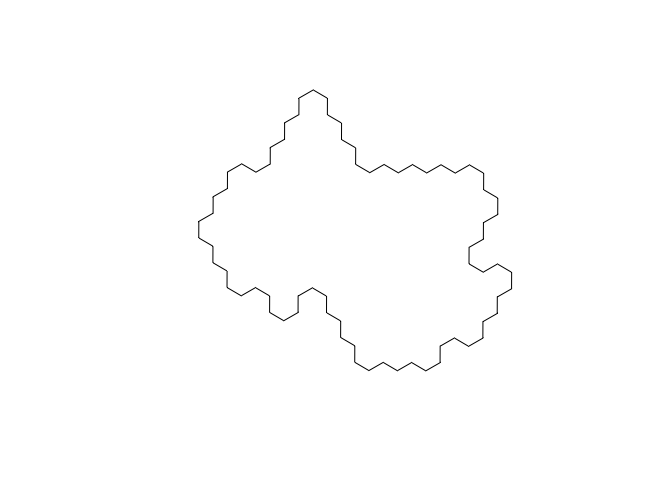
Next, we locate the centroid…
plot(st_union(hex))
plot(st_centroid(st_union(hex)), add=TRUE)
Finally, we pull out the coordinates of the centroid.
center <- st_coordinates(
st_centroid(
st_union(hex)
)
)
print(center)## X Y
## 1 -71.11408 42.38172These values can be passed to the setView function,
instead of hard-coding them.
map <- hex %>%
leaflet(
options = leafletOptions(
minZoom = 12
)
) %>%
setMaxBounds(
lng1 = bbox[['xmin']],
lat1 = bbox[['ymin']],
lng2 = bbox[['xmax']],
lat2 = bbox[['ymax']]
) %>%
# Access x and y or center here...
setView(center[,'X'], center[,'Y'], 13) %>%
addProviderTiles("Stamen.TonerLite") %>%
addPolygons(
fillColor = ~palette_quantiles(trees),
fillOpacity = 0.7,
opacity = 1,
weight = 1,
color='white',
highlight = highlightOptions(
weight = 3,
bringToFront = TRUE),
label = labels
) %>%
addLegend(
title = "Number of Street Trees",
pal = palette_quantiles,
opacity = 0.7,
values = trees,
labFormat = labelFormat(
digits = 0
)
)
map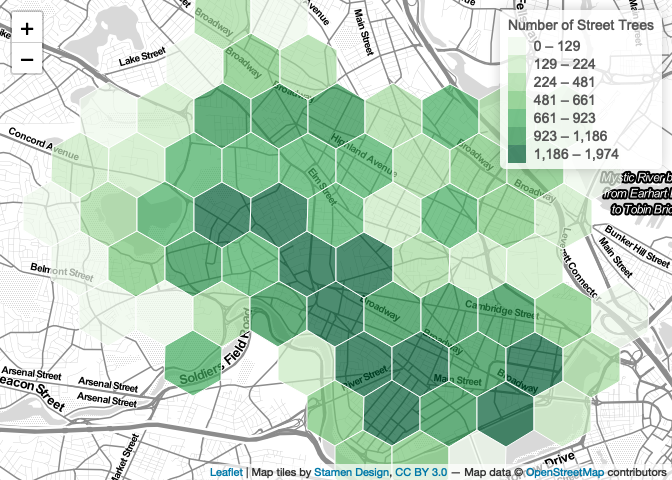
Exporting the Map
Once you’ve produced a map, you’ll probably want to export it. I do
this using the mapshot package, which allows you to quickly
and simply export a leaflet map object to either a ready-to-host HTML
file, or a image (.png, .jpg, etc.).
require('mapshot')## Loading required package: mapshot
## Warning in library(package, lib.loc = lib.loc, character.only = TRUE, logical.return = TRUE, : there is no package called 'mapshot'# Export to an HTML file...
mapshot(map, url = "street_trees.html")
# Export to a image file...
mapshot(map, file = "street_trees.png")When you first try to export your map, you may see this message:
PhantomJS not found. You can install it with webshot::install_phantomjs(). If it is installed, please make sure the phantomjs executable can be found via the PATH variable.If you do, simply run webshot::install_phantomjs() (as
directed)! Opening the html file in a web browser, you should see your
map, as designed in R, ready to be hosted and displayed publicly!
